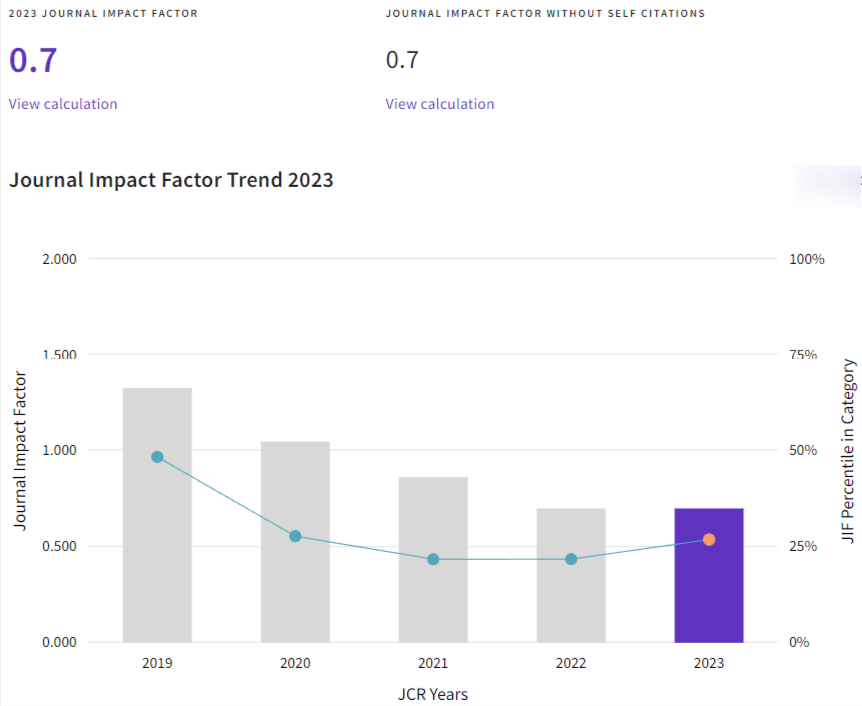
Análisis de QTL asociados a polimorfismos de nucleótido único (SNP) involucrados en el fenotipo lechero del ganado Holstein
DOI:
https://doi.org/10.22319/rmcp.v11i4.5295Palabras clave:
QTL, SNP, HolsteinResumen
El objetivo fue identificar QTL asociados a polimorfismos de nucleótido único (SNP) cuya acción contribuya al desarrollo fenotípico productivo, reproductivo y de salud del ganado lechero de raza Holstein. Ubicados en 120 genes del genoma Bos taurus UMD_3.1.1, se identificaron 341 QTL asociados a 189 SNP con efectos sobre rasgos productivos (FY, NM, MY, MTCAS, MBLF y PL), reproductivos (CONCRATE, DPR, EMBSUR, DAYOPEN y CONCEPT) y de salud (SCC, BTBS y RESRATE). Los SNP fueron verificados en la base de datos dbSNP-NCBI donde 42 % se ubicaron en intrones. Con la plataforma Jvenn se supo que los SNP rs135744058, rs110828053 y rs109503725, fueron comunes en los tres rasgos. La red de correlaciones entre rasgos y genes generada por MetScape (Cytoscape 3.4), mostró una correlación positiva entre PL, DPR, DAYOPEN, CONCRATE y CONCEPT; y una negativa de FY hacia PL, NM, DPR y CONCRATE. La funcionalidad de cada gen fue validada en las bases Gene-NCBI y UniProt, y por ClueGo (Cytoscape 3.4) se seleccionaron vías funcionales con un valor de significancia menor a 0.05, lo que evidenció un entrelazamiento entre el desarrollo de la glándula mamaria, la activación del sistema inmune y la respuesta a hormonas esteroideas; siendo el gen GH quien dirige dicha funcionalidad. Aunque el panorama genético muestra que existe un antagonismo entre rasgos productivos y reproductivos, la actividad genética funcional debido a los 189 SNP analizados, muestra una acción entrelazada en vías ontológicas que influyen tanto en los procesos de producción, así como en vías de reproductivas y de salud.
Descargas
Citas
Richardson IW, Berry DP, Wiencko HL, Higgins IM, More SJ, McClure J, et al. A genome-wide association study for genetic susceptibility to Mycobacterium bovis infection in dairy cattle identifies a susceptibility QTL on chromosome 23. Genet Sel Evol 2016;48:19-31.
Hu ZL, Park CA, Reecy JM. Developmental progress and current status of the Animal QTLdb. Nucleic Acids Res 2016;44(D1):D827-D833.
Yudin NS, Voevoda MI. Molecular genetic markers of economically important traits in dairy cattle. Russian J Genetics 2015;51(5):506-517.
Abo-Ismail MK, Brito LF, Miller SP, Sargolzaei M, Grossi DA, Moore SS, et al. Genome-wide association studies and genomic prediction of breeding values for calving performance and body conformation traits in Holstein cattle. Genet Sel Evol 2017;49(1):82-110.
Pegolo S, Mach N, Ramayo-Caldas Y, Schiavon S, Bittante G, Cecchinato A. Integration of GWAS, pathway and network analyses reveals novel mechanistic insights into the synthesis of milk proteins in dairy cows. Sci Rep 2018;8(1):566-580.
Wang M, Hancock TP, Chamberlain AJ, Vander Jagt CJ, Pryce JE, Cocks BG, et al. Putative bovine topological association domains and CTCF binding motifs can reduce the search space for causative regulatory variants of complex traits. BMC Genomics 2018;19(1):395-411.
Nayeri S, Sargolzaei M, Abo-Ismail MK, May N, Miller SP, Schenkel F, et al. Genome-wide association for milk production and female fertility traits in Canadian dairy Holstein cattle. BMC Genet 2016;17(1):75-85.
Cochran SD, Cole JB, Null DJ, Hansen PJ. Discovery of single nucleotide polymorphisms in candidate genes associated with fertility and production traits in Holstein cattle. BMC genetics 2013;14(1):49-71.
Bardou P, Mariette J, Escudié F, Djemiel C, Klopp C. jvenn: an interactive Venn diagram viewer. BMC bioinformatics 2014;15(1):293-299.
Basu S, Duren W, Evans CR, Burant CF, Michailidis G, Karnovsky A. Sparse network modeling and metscape-based visualization methods for the analysis of large-scale metabolomics data. Bioinformatics 2017;33(10):1545-1553.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome research 2003;13(11):2498-2504.
Duran Aguilar M, Roman Ponce SI, Ruiz Lopez FJ, Gonzalez Padilla E, Vasquez Pelaez CG, Bagnato A, et al. Genome-wide association study for milk somatic cell score in Holstein cattle using copy number variation as markers. J Anim Breed Genet 2017;134(1):49-59.
Karim L, Takeda H, Lin L, Druet T, Arias JA, Baurain D, et al. Variants modulating the expression of a chromosome domain encompassing PLAG1 influence bovine stature. Nat Genet 2011;43(5):405-413.
Kolbehdari D, Wang Z, Grant JR, Murdoch B, Prasad A, Xiu Z, et al. A whole genome scan to map QTL for milk production traits and somatic cell score in Canadian Holstein bulls. J Anim Breed Genet 2009;126(3):216-227.
Fang M, Fu W, Jiang D, Zhang Q, Sun D, Ding X, et al. A multiple-SNP approach for genome-wide association study of milk production traits in Chinese Holstein cattle. PLoS One 2014;9(8):e99544-e99551.
Gambra R, Penagaricano F, Kropp J, Khateeb K, Weigel KA, Lucey J, et al. Genomic architecture of bovine kappa-casein and beta-lactoglobulin. J Dairy Sci 2013;96(8):5333-5343.
Gervais O, Pong-Wong R, Navarro P, Haley CS, Nagamine Y. Antagonistic genetic correlations for milking traits within the genome of dairy cattle. PLoS One 2017;12(4):e0175105-e0175117.
Cole JB, Wiggans GR, Ma L, Sonstegard TS, Lawlor TJ, Crooker BA, et al. Genome-wide association analysis of thirty one production, health, reproduction and body conformation traits in contemporary US Holstein cows. BMC genomics 2011;12(1):408.
Pesek P, Pribyl J, Vostry L. Genetic variances of SNP loci for milk yield in dairy cattle. J Appl Genet 2015;56(3):339-347.
Pimentel EC, Bauersachs S, Tietze M, Simianer H, Tetens J, Thaller G, et al. Exploration of relationships between production and fertility traits in dairy cattle via association studies of SNPs within candidate genes derived by expression profiling. Anim Genet 2011;42(3):251-262.
Huang W, Penagaricano F, Ahmad KR, Lucey JA, Weigel KA, Khatib H. Association between milk protein gene variants and protein composition traits in dairy cattle. J Dairy Sci 2012;95(1):440-449.
Schopen GC, Visker MH, Koks PD, Mullaart E, van Arendonk JA, Bovenhuis H. Whole-genome association study for milk protein composition in dairy cattle. J Dairy Sci 2011;94(6):3148-158.
Viale E, Tiezzi F, Maretto F, De Marchi M, Penasa M, Cassandro M. Association of candidate gene polymorphisms with milk technological traits, yield, composition, and somatic cell score in Italian Holstein-Friesian sires. J Dairy Sci 2017;100(9):7271-7281.
Fontanesi L, Calo DG, Galimberti G, Negrini R, Marino R, Nardone A, et al. A candidate gene association study for nine economically important traits in Italian Holstein cattle. Anim Genet 2014;45(4):576-580.
Wang W, Cheng L, Yi J, Gan J, Tang H, Fu MZ, et al. Health and production traits in bovine are associated with single nucleotide polymorphisms in the NOD2 gene. Genet Mol Res 2015;14(2):3570-3578.
Finlay EK, Berry DP, Wickham B, Gormley EP, Bradley DG. A genome wide association scan of bovine tuberculosis susceptibility in Holstein-Friesian dairy cattle. PLoS One 2012;7(2):e30545-e135094.
Wang Y, Wang S, Liu T, Tu W, Li W, Dong G, et al. CARD15 Gene Polymorphisms Are Associated with Tuberculosis Susceptibility in Chinese Holstein Cows. PLoS One 2015;10(8):e0135085.
Ortega MS, Denicol AC, Cole JB, Null DJ, Hansen PJ. Use of single nucleotide polymorphisms in candidate genes associated with daughter pregnancy rate for prediction of genetic merit for reproduction in Holstein cows. Anim Genetics 2016;47(3):288-297.
Dikmen S, Wang XZ, Ortega MS, Cole JB, Null DJ, Hansen PJ. Single nucleotide polymorphisms associated with thermoregulation in lactating dairy cows exposed to heat stress. J Anim Breed Genet 2015;132(6):409-419.
Srivastava U, Aromolaran AS, Fabris F, Lazaro D, Kassotis J, Qu Y, et al. Novel function of alpha1D L-type calcium channel in the atria. Biochem Biophys Res Commun 2017;482(4):771-776.
Peruffo A, Giacomello M, Montelli S, Panin M, Cozzi B. Expression profile of the pore-forming subunits alpha1A and alpha1D in the foetal bovine hypothalamus: a mammal with a long gestation. Neurosci Lett 2013;556:124-128.
Cerny KL, Garrett E, Walton AJ, Anderson LH, Bridges PJ. A transcriptomal analysis of bovine oviductal epithelial cells collected during the follicular phase versus the luteal phase of the estrous cycle. Reprod Biol Endocrinol 2015;13:84-96.
Kemilainen H, Adam M, Maki-Jouppila J, Damdimopoulou P, Damdimopoulos AE, Kere J, et al. The hydroxysteroid (17beta) dehydrogenase family gene HSD17B12 is involved in the prostaglandin synthesis pathway, the ovarian function, and regulation of fertility. Endocrinology 2016;157(10):3719-3730.
Gehmlich K, Lambiase PD, Asimaki A, Ciaccio EJ, Ehler E, Syrris P, et al. A novel desmocollin-2 mutation reveals insights into the molecular link between desmosomes and gap junctions. Heart Rhythm 2011;8(5):711-718.
Pryce J, Royal M, Garnsworthy P, Mao IL. Fertility in the high-producing dairy cow. Livestock Prod Sci 2004;86(1-3):125-135.
Pryce J, Veerkamp R. The incorporation of fertility indices in genetic improvement programmes. BSAP Occasional Publication 2001;26(1):237-249.
Peñagaricano F, Khatib H. Association of milk protein genes with fertilization rate and early embryonic development in Holstein dairy cattle. J Dairy Res 2011;79(01):47-52.
Cushman R, DeSouza J, Hedgpeth V, Britt J. Effect of long-term treatment with recombinant bovine somatotropin and estradiol on hormone concentrations and ovulatory response of superovulated cattle. Theriogenology 2001;55(7):1533-1547.
Akers RM. A 100-Year Review: Mammary development and lactation. J Dairy Sci 2017;100(12):10332-10352.
Descargas
Publicado
Cómo citar
-
Resumen724
-
PDF360
-
PDF 178
-
Texto completo56
Número
Sección
Licencia
Los autores/as que publiquen en la Revista Mexicana de Ciencias Pecuarias aceptan las siguientes condiciones:
De acuerdo con la legislación de derechos de autor, la Revista Mexicana de Ciencias Pecuarias reconoce y respeta el derecho moral de los autores/as, así como la titularidad del derecho patrimonial, el cual será cedido a la revista para su difusión en acceso abierto.

Esta obra está bajo una Licencia Creative Commons Atribución-NoComercial-CompartirIgual 4.0 Internacional.